ARG42493
anti-Histone H2A.X phospho (Ser139) antibody
anti-Histone H2A.X phospho (Ser139) antibody for ICC/IF,IHC-Formalin-fixed paraffin-embedded sections,Western blot and Human,Mouse,Rat
Overview
| Product Description | Rabbit Polyclonal antibody recognizes Histone H2A.X phospho (Ser139) |
|---|---|
| Tested Reactivity | Hu, Ms, Rat |
| Tested Application | ICC/IF, IHC-P, WB |
| Host | Rabbit |
| Clonality | Polyclonal |
| Isotype | IgG |
| Target Name | Histone H2A.X |
| Antigen Species | Human |
| Immunogen | Phosphospecific peptide around Ser139 of Human Histone H2A.X (NP_002096.1). |
| Conjugation | Un-conjugated |
| Alternate Names | H2AX; H2a/x; H2A.X; Histone H2AX; H2A/X; Histone H2A.X; gamma H2AX; gamma H2A.X |
Application Instructions
| Application Suggestion |
|
||||||||
|---|---|---|---|---|---|---|---|---|---|
| Application Note | * The dilutions indicate recommended starting dilutions and the optimal dilutions or concentrations should be determined by the scientist. | ||||||||
| Positive Control | Jurkat + Etoposide NIH/3T3 + UV C6 + UV |
||||||||
| Observed Size | ~ 17 kDa |
Properties
| Form | Liquid |
|---|---|
| Purification | Affinity purification with immunogen. |
| Buffer | PBS (pH 7.3), 0.02% Sodium azide and 50% Glycerol. |
| Preservative | 0.02% Sodium azide |
| Stabilizer | 50% Glycerol |
| Storage Instruction | For continuous use, store undiluted antibody at 2-8°C for up to a week. For long-term storage, aliquot and store at -20°C. Storage in frost free freezers is not recommended. Avoid repeated freeze/thaw cycles. Suggest spin the vial prior to opening. The antibody solution should be gently mixed before use. |
| Note | For laboratory research only, not for drug, diagnostic or other use. |
Bioinformation
| Database Links | |
|---|---|
| Gene Symbol | H2AFX |
| Gene Full Name | H2A histone family, member X |
| Background | Histones are basic nuclear proteins that are responsible for the nucleosome structure of the chromosomal fiber in eukaryotes. Two molecules of each of the four core histones (H2A, H2B, H3, and H4) form an octamer, around which approximately 146 bp of DNA is wrapped in repeating units, called nucleosomes. The linker histone, H1, interacts with linker DNA between nucleosomes and functions in the compaction of chromatin into higher order structures. This gene encodes a replication-independent histone that is a member of the histone H2A family, and generates two transcripts through the use of the conserved stem-loop termination motif, and the polyA addition motif. [provided by RefSeq, Oct 2015] |
| Function | Variant histone H2A which replaces conventional H2A in a subset of nucleosomes. Nucleosomes wrap and compact DNA into chromatin, limiting DNA accessibility to the cellular machineries which require DNA as a template. Histones thereby play a central role in transcription regulation, DNA repair, DNA replication and chromosomal stability. DNA accessibility is regulated via a complex set of post-translational modifications of histones, also called histone code, and nucleosome remodeling. Required for checkpoint-mediated arrest of cell cycle progression in response to low doses of ionizing radiation and for efficient repair of DNA double strand breaks (DSBs) specifically when modified by C-terminal phosphorylation. [UniProt] |
| Cellular Localization | Chromosome. [UniProt] |
| Highlight | Related products: Histone H2A.X antibodies; Anti-Rabbit IgG secondary antibodies; Related news: Senescence Marker Antibody Panel is launched |
| Calculated MW | 15 kDa |
| PTM | Phosphorylated on Ser-140 (to form gamma-H2AX or H2AX139ph) in response to DNA double strand breaks (DSBs) generated by exogenous genotoxic agents and by stalled replication forks, and may also occur during meiotic recombination events and immunoglobulin class switching in lymphocytes. Phosphorylation can extend up to several thousand nucleosomes from the actual site of the DSB and may mark the surrounding chromatin for recruitment of proteins required for DNA damage signaling and repair. Widespread phosphorylation may also serve to amplify the damage signal or aid repair of persistent lesions. Phosphorylation of Ser-140 (H2AX139ph) in response to ionizing radiation is mediated by both ATM and PRKDC while defects in DNA replication induce Ser-140 phosphorylation (H2AX139ph) subsequent to activation of ATR and PRKDC. Dephosphorylation of Ser-140 by PP2A is required for DNA DSB repair. In meiosis, Ser-140 phosphorylation (H2AX139ph) may occur at synaptonemal complexes during leptotene as an ATM-dependent response to the formation of programmed DSBs by SPO11. Ser-140 phosphorylation (H2AX139ph) may subsequently occurs at unsynapsed regions of both autosomes and the XY bivalent during zygotene, downstream of ATR and BRCA1 activation. Ser-140 phosphorylation (H2AX139ph) may also be required for transcriptional repression of unsynapsed chromatin and meiotic sex chromosome inactivation (MSCI), whereby the X and Y chromosomes condense in pachytene to form the heterochromatic XY-body. During immunoglobulin class switch recombination in lymphocytes, Ser-140 phosphorylation (H2AX139ph) may occur at sites of DNA-recombination subsequent to activation of the activation-induced cytidine deaminase AICDA. Phosphorylation at Tyr-143 (H2AXY142ph) by BAZ1B/WSTF determines the relative recruitment of either DNA repair or pro-apoptotic factors. Phosphorylation at Tyr-143 (H2AXY142ph) favors the recruitment of APBB1/FE65 and pro-apoptosis factors such as MAPK8/JNK1, triggering apoptosis. In contrast, dephosphorylation of Tyr-143 by EYA proteins (EYA1, EYA2, EYA3 or EYA4) favors the recruitment of MDC1-containing DNA repair complexes to the tail of phosphorylated Ser-140 (H2AX139ph). Monoubiquitination of Lys-120 (H2AXK119ub) by RING1 and RNF2/RING2 complex gives a specific tag for epigenetic transcriptional repression (By similarity). Following DNA double-strand breaks (DSBs), it is ubiquitinated through 'Lys-63' linkage of ubiquitin moieties by the E2 ligase UBE2N and the E3 ligases RNF8 and RNF168, leading to the recruitment of repair proteins to sites of DNA damage. Ubiquitination at Lys-14 and Lys-16 (H2AK13Ub and H2AK15Ub, respectively) in response to DNA damage is initiated by RNF168 that mediates monoubiquitination at these 2 sites, and 'Lys-63'-linked ubiquitin are then conjugated to monoubiquitin; RNF8 is able to extend 'Lys-63'-linked ubiquitin chains in vitro. H2AK119Ub and ionizing radiation-induced 'Lys-63'-linked ubiquitination (H2AK13Ub and H2AK15Ub) are distinct events. Acetylation at Lys-37 increases in S and G2 phases. This modification has been proposed to play a role in DNA double-strand break repair (By similarity). [UniProt] |
Images (4) Click the Picture to Zoom In
-
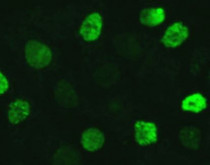
ARG42493 anti-Histone H2A.X phospho (Ser139) antibody ICC/IF image
Immunofluorescence: HeLa cells were treated by UV for 15-30 minutes at RT. Cells were stained with ARG42493 anti-Histone H2A.X phospho (Ser139) antibody at 1:100 dilution.
-
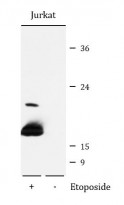
ARG42493 anti-Histone H2A.X phospho (Ser139) antibody WB image
Western blot: Jurkat cells were untreated (right) or treated by Etoposide (25 µM) for 5 hours (left). 25 µg of cell lysates were stained with ARG42493 anti-Histone H2A.X phospho (Ser139) antibody at 1:1000 dilution.
-
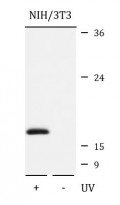
ARG42493 anti-Histone H2A.X phospho (Ser139) antibody WB image
Western blot: NIH/3T3 cells were untreated (right) or treated by UV for 15-30 minutes (left). 25 µg of cell lysates were stained with ARG42493 anti-Histone H2A.X phospho (Ser139) antibody at 1:1000 dilution.
-
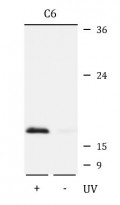
ARG42493 anti-Histone H2A.X phospho (Ser139) antibody WB image
Western blot: C6 cells were untreated (right) or treated by UV for 15-30 minutes (left). 25 µg of cell lysates were stained with ARG42493 anti-Histone H2A.X phospho (Ser139) antibody at 1:1000 dilution.